Bayesian Function-on-Function Regression (FoFR)
2026-04-06
Source:vignettes/fofr_bayes_vignette.Rmd
fofr_bayes_vignette.RmdIntroduction
This vignette provides a detailed guide to the
fofr_bayes() function in the refundBayes
package, which fits Bayesian Function-on-Function Regression (FoFR)
models using Stan. FoFR extends both the scalar-on-function regression
(SoFR) and function-on-scalar regression (FoSR) frameworks by modeling a
functional response as a function of one or more
functional predictors, along with optional scalar
covariates.
In contrast to SoFR, where the outcome is scalar and the predictors
are functional, and to FoSR, where the outcome is functional and the
predictors are scalar, FoFR allows the effect of a functional predictor
on a functional response to be captured through a bivariate
coefficient function
.
The model is specified using the same mgcv-style syntax as
the other regression functions in the refundBayes
package.
The methodology extends the framework described in Jiang, Crainiceanu, and Cui (2025), Tutorial on Bayesian Functional Regression Using Stan, published in Statistics in Medicine.
Install the refundBayes Package
The refundBayes package can be installed from CRAN:
install.packages("refundBayes")For the latest version of the refundBayes package, users
can install from GitHub:
library(remotes)
remotes::install_github("https://github.com/ZirenJiang/refundBayes")Statistical Model
The FoFR Model
Function-on-Function Regression (FoFR) models the relationship between a functional response and one or more functional predictors, optionally adjusted for scalar covariates. Both the response and the predictors are curves observed over (potentially different) continua.
For subject , let be the functional response observed at time points over a response domain , and let for be a functional predictor observed at points over a predictor domain . Let be a vector of scalar predictors (the first covariate may be an intercept, ). The FoFR model assumes:
where:
- are functional coefficients for the scalar predictors, each describing how the -th scalar predictor affects the response at each point ,
- is the bivariate coefficient function that characterizes how the functional predictor at predictor-domain location affects the response at response-domain location ,
- is the residual process, which is correlated across .
The integral is approximated using a Riemann sum over the observed predictor-domain grid points. The domains and are not restricted to ; they are determined by the actual observation grids in the data.
When multiple functional predictors are present, the model extends naturally:
where is the bivariate coefficient function for the -th functional predictor.
Modeling the Residual Structure
To account for the within-subject correlation in the residuals, the model decomposes using functional principal components, following the same approach as in FoSR (Goldsmith, Zipunnikov, and Schrack, 2015):
where
are the eigenfunctions estimated via FPCA (using
refund::fpca.face),
are the subject-specific FPCA scores with
,
and
is independent measurement error. The eigenvalues
and the error variance
are estimated from the data.
Scalar Predictor Coefficients via Penalized Splines
Each scalar predictor coefficient function is represented using spline basis functions in the response domain:
Smoothness is induced through a quadratic penalty:
where is the penalty matrix derived from the spline basis and is the smoothing parameter for the -th scalar predictor, estimated from the data.
Bivariate Coefficient via Tensor Product Basis
The key feature of FoFR is the bivariate coefficient function , which lives on the product domain . This function is represented using a tensor product of two sets of basis functions:
Predictor-domain basis: , constructed from the
s()term in the formula using themgcvspline basis (e.g., cubic regression splines). These are further decomposed into random and fixed effect components via the spectral reparametrisation ofmgcv::smooth2random(), yielding (random effects) and (fixed effects).Response-domain basis: , the same spline basis used for the scalar predictor coefficients.
The bivariate coefficient is then:
where is the matrix of random effect coefficients and is the matrix of fixed effect coefficients.
With this representation, the integral contribution of the functional predictor becomes:
where and are the and row vectors of the transformed predictor-domain design matrices for subject (the same matrices used in SoFR).
In matrix notation for all subjects, the functional predictor contribution to the mean is:
which is an matrix, where is the matrix of response-domain spline basis evaluations.
Dual-Direction Smoothness Penalties
Because the bivariate coefficient lives on a two-dimensional domain, smoothness must be enforced in both directions:
Predictor-domain smoothness (-direction)
Following the same approach as in SoFR, the random effect coefficients use a variance-component reparametrisation:
where is the predictor-domain smoothing parameter. This is the standard non-centered parameterisation that separates the scale () from the direction (), improving sampling efficiency in Stan.
Response-domain smoothness (-direction)
The response-domain penalty matrix is applied row-wise to the coefficient matrices. For each row of and :
where is the response-domain smoothing parameter. This ensures that is smooth in the -direction for every fixed .
The two smoothing parameters and are estimated from the data with weakly informative inverse-Gamma priors.
Full Bayesian Model
The complete Bayesian FoFR model combines the mean structure, residual FPCA decomposition, and all priors:
In matrix form for all subjects:
where is the matrix of functional responses, is the scalar design matrix, is the matrix of scalar predictor spline coefficients, is the matrix of FPCA scores, and is the matrix of eigenfunctions.
Prior Specification
The full prior specification is:
| Parameter | Prior | Description |
|---|---|---|
| (scalar predictor spline coefs) | Penalized spline prior for smoothness of | |
| (scalar smoothing parameter) | Weakly informative prior on smoothing | |
| (standardized random effects) | Non-centered parameterisation for -direction | |
| (predictor-domain smoothing) | Prior on predictor-domain smoothing | |
| (row-wise) | Penalized spline prior for response-domain smoothness | |
| (response-domain smoothing) | Prior on response-domain smoothing | |
| (FPCA scores) | FPCA score distribution | |
| (FPCA eigenvalues) | Weakly informative prior on eigenvalues | |
| (residual variance) | Weakly informative prior on noise variance |
Optional: Joint FPCA Modeling
The default fofr_bayes() model treats the observed
functional predictor
as if it were observed without error. When the predictor curves are
noisy, this attenuates the bivariate coefficient
toward zero (errors-in-variables bias) and shrinks the credible-band
width. The joint_FPCA argument activates an alternative
model in which
is replaced by an FPCA representation and the subject-specific FPC
scores are sampled jointly with the bivariate
coefficient, propagating the FPCA uncertainty into the posterior of
.
The same joint_FPCA argument is available in
sofr_bayes() and fcox_bayes(). The full model
specification (FPCA likelihood for the predictor, FPC-score prior
centered on the initial refund::fpca.sc() scores, the
resulting Stan program with bivariate coefficients in the FPC basis, and
a worked FoFR example) is presented in the dedicated Joint FPCA vignette.
Relationship to SoFR and FoSR
The FoFR model nests both the SoFR and FoSR models as special cases:
| Model | Response | Predictors | Coefficient | Implemented in |
|---|---|---|---|---|
| SoFR | Scalar | Functional | Univariate | sofr_bayes() |
| FoSR | Functional | Scalar | Univariate | fosr_bayes() |
| FoFR | Functional | Functional + Scalar | Bivariate + Univariate | fofr_bayes() |
The fofr_bayes() function inherits:
- From FoSR: the response-domain spline basis , the FPCA residual structure, and the scalar predictor handling.
- From SoFR: the predictor-domain spectral reparametrisation
(random/fixed effect decomposition via
mgcv::smooth2random()), the basis extraction, and the functional coefficient reconstruction.
The fofr_bayes() Function
Usage
fofr_bayes(
formula,
data,
joint_FPCA = NULL,
runStan = TRUE,
niter = 3000,
nwarmup = 1000,
nchain = 3,
ncores = 1,
spline_type = "bs",
spline_df = 10
)Arguments
| Argument | Description |
|---|---|
formula |
Functional regression formula, using the same syntax as
mgcv::gam. The left-hand side is the functional response
(an
matrix in data). The right-hand side includes scalar
predictors as standard terms and functional predictors via
s(..., by = ...) terms. At least one functional predictor
must be present; otherwise use fosr_bayes(). |
data |
A data frame containing all variables used in the model. The functional response and functional predictor should be stored as and matrices, respectively. |
joint_FPCA |
A logical (TRUE/FALSE) vector
of the same length as the number of functional predictors, indicating
whether to jointly model FPCA for each functional predictor. When
TRUE, the observed functional predictor is replaced by an
FPCA representation and its FPC scores are sampled jointly with the
bivariate coefficient (errors-in-variables-aware fit). See the Joint FPCA vignette for the model
specification and a worked FoFR example. Default is NULL,
which sets all entries to FALSE (no joint FPCA). |
runStan |
Logical. Whether to run the Stan program. If
FALSE, the function only generates the Stan code and data
without sampling. This is useful for inspecting or modifying the
generated Stan code. Default is TRUE. |
niter |
Total number of Bayesian posterior sampling iterations
(including warmup). Default is 3000. |
nwarmup |
Number of warmup (burn-in) iterations. These samples
are discarded and not used for inference. Default is
1000. |
nchain |
Number of Markov chains for posterior sampling.
Multiple chains help assess convergence. Default is 3. |
ncores |
Number of CPU cores to use when executing the chains in
parallel. Default is 1. |
spline_type |
Type of spline basis used for the response-domain
component. Default is "bs" (B-splines). Other types
supported by mgcv may also be used. |
spline_df |
Number of degrees of freedom (basis functions) for the
response-domain spline basis. Default is 10. |
Return Value
The function returns a list of class "refundBayes"
containing the following elements:
| Element | Description |
|---|---|
stanfit |
The Stan fit object (class stanfit). Can
be used for convergence diagnostics, traceplots, and additional
summaries via the rstan package. |
spline_basis |
Basis functions used to reconstruct the functional coefficients from the posterior samples. |
stancode |
A character string containing the generated Stan model code. |
standata |
A list containing the data passed to the Stan model. |
scalar_func_coef |
A 3-d array
()
of posterior samples for scalar predictor coefficient functions
,
where
is the number of posterior samples,
is the number of scalar predictors, and
is the number of response-domain time points. NULL if no
scalar predictors. |
bivar_func_coef |
A list of 3-d arrays. Each element corresponds to one functional predictor and is an array of dimension , representing posterior samples of the bivariate coefficient function evaluated on the predictor-domain grid ( points) and response-domain grid ( points). |
func_coef |
Same as scalar_func_coef; included for
compatibility with the plot.refundBayes() method. |
family |
The model family: "fofr". |
Formula Syntax
The formula combines the FoSR syntax (functional response on the
left-hand side) with the SoFR syntax (functional predictors via
s() terms on the right-hand side):
Y_mat ~ X1 + X2 + s(sindex, by = X_func, bs = "cr", k = 10)where:
-
Y_mat: the name of the functional response variable indata. This should be an matrix, where each row contains the functional observations for one subject across response-domain time points. -
X1,X2: scalar predictor(s), included using standard formula syntax. -
s(sindex, by = X_func, bs = "cr", k = 10): the functional predictor term:-
sindex: an matrix of predictor-domain grid points. Each row contains the same observation points (replicated across subjects). -
X_func: an matrix of functional predictor values. The -th row contains the observed values for subject . -
bs: the type of spline basis for the predictor domain (e.g.,"cr"for cubic regression splines). -
k: the number of basis functions in the predictor domain.
-
The response-domain spline basis is controlled separately via the
spline_type and spline_df arguments to
fofr_bayes(). This design separates the two basis
specifications: the predictor-domain basis is specified in the formula
(as in SoFR), while the response-domain basis is specified via function
arguments (as in FoSR).
Multiple functional predictors can be included by adding additional
s() terms.
Example: Bayesian FoFR with Simulated Data
We demonstrate the fofr_bayes() function using a
simulation study with a known bivariate coefficient function
and a scalar predictor coefficient function
.
Simulate Data
library(refundBayes)
set.seed(42)
# --- Dimensions ---
n <- 200 # number of subjects
L <- 30 # number of predictor-domain grid points
M <- 30 # number of response-domain grid points
sindex <- seq(0, 1, length.out = L) # predictor domain grid
tindex <- seq(0, 1, length.out = M) # response domain grid
# --- Functional predictor X(s): smooth random curves ---
X_func <- matrix(0, nrow = n, ncol = L)
for (i in 1:n) {
X_func[i, ] <- rnorm(1) * sin(2 * pi * sindex) +
rnorm(1) * cos(2 * pi * sindex) +
rnorm(1) * sin(4 * pi * sindex) +
rnorm(1, sd = 0.3)
}
# --- Scalar predictor ---
age <- rnorm(n)
# --- True coefficient functions ---
# Bivariate coefficient: beta(s, t) = sin(2*pi*s) * cos(2*pi*t)
beta_true <- outer(sin(2 * pi * sindex), cos(2 * pi * tindex))
# Scalar coefficient function: alpha(t) = 0.5 * sin(pi*t)
alpha_true <- 0.5 * sin(pi * tindex)
# --- Generate functional response ---
# Y_i(t) = age_i * alpha(t) + integral X_i(s) beta(s,t) ds + epsilon_i(t)
signal_scalar <- outer(age, alpha_true) # n x M
signal_func <- (X_func %*% beta_true) / L # n x M (Riemann sum)
epsilon <- matrix(rnorm(n * M, sd = 0.3), nrow = n) # n x M
Y_mat <- signal_scalar + signal_func + epsilon
# --- Organize data ---
dat <- data.frame(age = age)
dat$Y_mat <- Y_mat
dat$X_func <- X_func
dat$sindex <- matrix(rep(sindex, n), nrow = n, byrow = TRUE)The simulated dataset dat contains:
-
Y_mat: an matrix of functional response values, -
age: a scalar predictor, -
X_func: an matrix of functional predictor values (smooth random curves), -
sindex: an matrix of predictor-domain grid points (identical rows).
The true data-generating model is:
Fit the Bayesian FoFR Model
fit_fofr <- fofr_bayes(
formula = Y_mat ~ age + s(sindex, by = X_func, bs = "cr", k = 10),
data = dat,
spline_type = "bs",
spline_df = 10,
niter = 2000,
nwarmup = 1000,
nchain = 3,
ncores = 3
)In this call:
- The formula specifies the functional response
Y_mat(an matrix) with one scalar predictorageand one functional predictorX_func. - The predictor-domain spline basis uses cubic regression splines
(
bs = "cr") withk = 10basis functions. - The response-domain spline basis uses B-splines
(
spline_type = "bs") withspline_df = 10degrees of freedom. - The eigenfunctions for the residual structure are estimated
automatically via FPCA using
refund::fpca.face. - The sampler runs 3 chains in parallel, each with 2000 total iterations (1000 warmup + 1000 posterior samples).
A Note on Computation
FoFR models are the most computationally demanding among the models
in refundBayes because the Stan program estimates bivariate
coefficient matrices (with
parameters per functional predictor) in addition to the scalar predictor
coefficients and FPCA scores. For exploratory analyses, consider using
fewer basis functions (e.g., k = 5,
spline_df = 5) and a single chain. For final inference, use
the full setup with multiple chains and convergence diagnostics.
Visualisation
Bivariate Coefficient
The estimated bivariate coefficient
is stored as a 3-d array in bivar_func_coef. The posterior
mean surface and comparison with the truth can be visualised using
heatmaps:
# Posterior mean of the bivariate coefficient
beta_est <- apply(fit_fofr$bivar_func_coef[[1]], c(2, 3), mean)
# Pointwise 95% credible interval bounds
beta_lower <- apply(fit_fofr$bivar_func_coef[[1]], c(2, 3),
function(x) quantile(x, 0.025))
beta_upper <- apply(fit_fofr$bivar_func_coef[[1]], c(2, 3),
function(x) quantile(x, 0.975))
# Side-by-side heatmaps: true vs estimated vs difference
par(mfrow = c(1, 3), mar = c(4, 4, 2, 1))
image(sindex, tindex, beta_true,
xlab = "s (predictor domain)", ylab = "t (response domain)",
main = expression("True " * beta(s, t)),
col = hcl.colors(64, "Blue-Red 3"))
image(sindex, tindex, beta_est,
xlab = "s (predictor domain)", ylab = "t (response domain)",
main = expression("Estimated " * hat(beta)(s, t)),
col = hcl.colors(64, "Blue-Red 3"))
image(sindex, tindex, beta_est - beta_true,
xlab = "s (predictor domain)", ylab = "t (response domain)",
main = "Difference (Est - True)",
col = hcl.colors(64, "Blue-Red 3"))For richer 3-d surface visualisations, use
fields::image.plot() or plotly::plot_ly() with
type "surface".
Scalar Coefficient Function
The estimated scalar predictor coefficient function can be plotted with pointwise credible intervals:
alpha_est <- apply(fit_fofr$scalar_func_coef[, 1, ], 2, mean)
alpha_lower <- apply(fit_fofr$scalar_func_coef[, 1, ], 2,
function(x) quantile(x, 0.025))
alpha_upper <- apply(fit_fofr$scalar_func_coef[, 1, ], 2,
function(x) quantile(x, 0.975))
par(mfrow = c(1, 1))
plot(tindex, alpha_true, type = "l", lwd = 2, col = "black",
ylim = range(c(alpha_lower, alpha_upper)),
xlab = "t (response domain)", ylab = expression(alpha(t)),
main = "Scalar coefficient function: age")
lines(tindex, alpha_est, col = "blue", lwd = 2)
polygon(c(tindex, rev(tindex)),
c(alpha_lower, rev(alpha_upper)),
col = rgb(0, 0, 1, 0.2), border = NA)
legend("topright",
legend = c("Truth", "Posterior mean", "95% CI"),
col = c("black", "blue", rgb(0, 0, 1, 0.2)),
lwd = c(2, 2, 10), bty = "n")Slices of the Bivariate Coefficient
To examine at fixed values of or , extract slices from the posterior:
# Fix s at the midpoint of the predictor domain and plot beta(s_mid, t)
s_mid_idx <- which.min(abs(sindex - 0.5))
beta_slice_est <- apply(fit_fofr$bivar_func_coef[[1]][, s_mid_idx, ], 2, mean)
beta_slice_lower <- apply(fit_fofr$bivar_func_coef[[1]][, s_mid_idx, ], 2,
function(x) quantile(x, 0.025))
beta_slice_upper <- apply(fit_fofr$bivar_func_coef[[1]][, s_mid_idx, ], 2,
function(x) quantile(x, 0.975))
beta_slice_true <- beta_true[s_mid_idx, ]
plot(tindex, beta_slice_true, type = "l", lwd = 2, col = "black",
ylim = range(c(beta_slice_lower, beta_slice_upper)),
xlab = "t (response domain)",
ylab = expression(beta(s[mid], t)),
main = paste0("Slice at s = ", round(sindex[s_mid_idx], 2)))
lines(tindex, beta_slice_est, col = "red", lwd = 2)
polygon(c(tindex, rev(tindex)),
c(beta_slice_lower, rev(beta_slice_upper)),
col = rgb(1, 0, 0, 0.2), border = NA)
legend("topright",
legend = c("Truth", "Posterior mean", "95% CI"),
col = c("black", "red", rgb(1, 0, 0, 0.2)),
lwd = c(2, 2, 10), bty = "n")Inspecting the Generated Stan Code
Setting runStan = FALSE allows you to inspect or modify
the Stan code before running the model:
# Generate Stan code without running the sampler
fofr_code <- fofr_bayes(
formula = Y_mat ~ age + s(sindex, by = X_func, bs = "cr", k = 10),
data = dat,
spline_type = "bs",
spline_df = 10,
runStan = FALSE
)
# Print the generated Stan code
cat(fofr_code$stancode)The generated Stan code includes all five standard blocks
(data, transformed data,
parameters, transformed parameters,
model). The parameters block declares
matrix-valued parameters for the bivariate coefficients, and the
model block includes both
-direction
and
-direction
smoothness priors.
Simulation Study: Bayesian vs Frequentist FoFR
To benchmark fofr_bayes() against the standard
frequentist function-on-function regression fit, we ran a simulation
study comparing posterior-mean prediction against the penalised
tensor-product estimator implemented in refund::pffr(). The
full simulation script is shipped as
Simulation/FoFR_Simulation_V3.R, with a stand-alone Stan
program in Simulation/StanFoFR_Gaussian.stan.
Simulation Setup
Functional observations were generated from a function-on-function model without scalar predictors:
on uniform grids of size over . Functional predictors were generated as four-component Fourier expansions with eigenvalues . Two true bivariate-coefficient surfaces and three signal-strength levels were considered:
| Factor | Levels |
|---|---|
| Sample size | |
| -surface type | type 1: separable ; type 2: Gaussian bump |
| Signal-strength multiplier |
Each cell of the design was replicated approximately 500 times, giving roughly 9000 fits per method.
Comparator Methods
-
Frequentist baseline –
refund::pffr()with a singleff(X_func, xind = sgrid, integration = "riemann", splinepars = list(bs = "ps", k = c(10, 10)))term and the response indexed byyind = tgrid. Theintegration = "riemann"option is critical here: the data-generating quadrature is left-Riemann with uniform weight , and matching this inpffr()(rather than the default Simpson rule) keeps on the same pointwise scale as during recovery comparisons. -
Bayesian model –
refundBayes::fofr_bayes()(or the equivalent standalone Stan program inSimulation/StanFoFR_Gaussian.stan), withk = 10predictor-domain cubic-regression splines,spline_df = 10response-domain B-splines, FPCA residual structure estimated viarefund::fpca.sc(), 1 chain, 1000 iterations (500 warm-up).
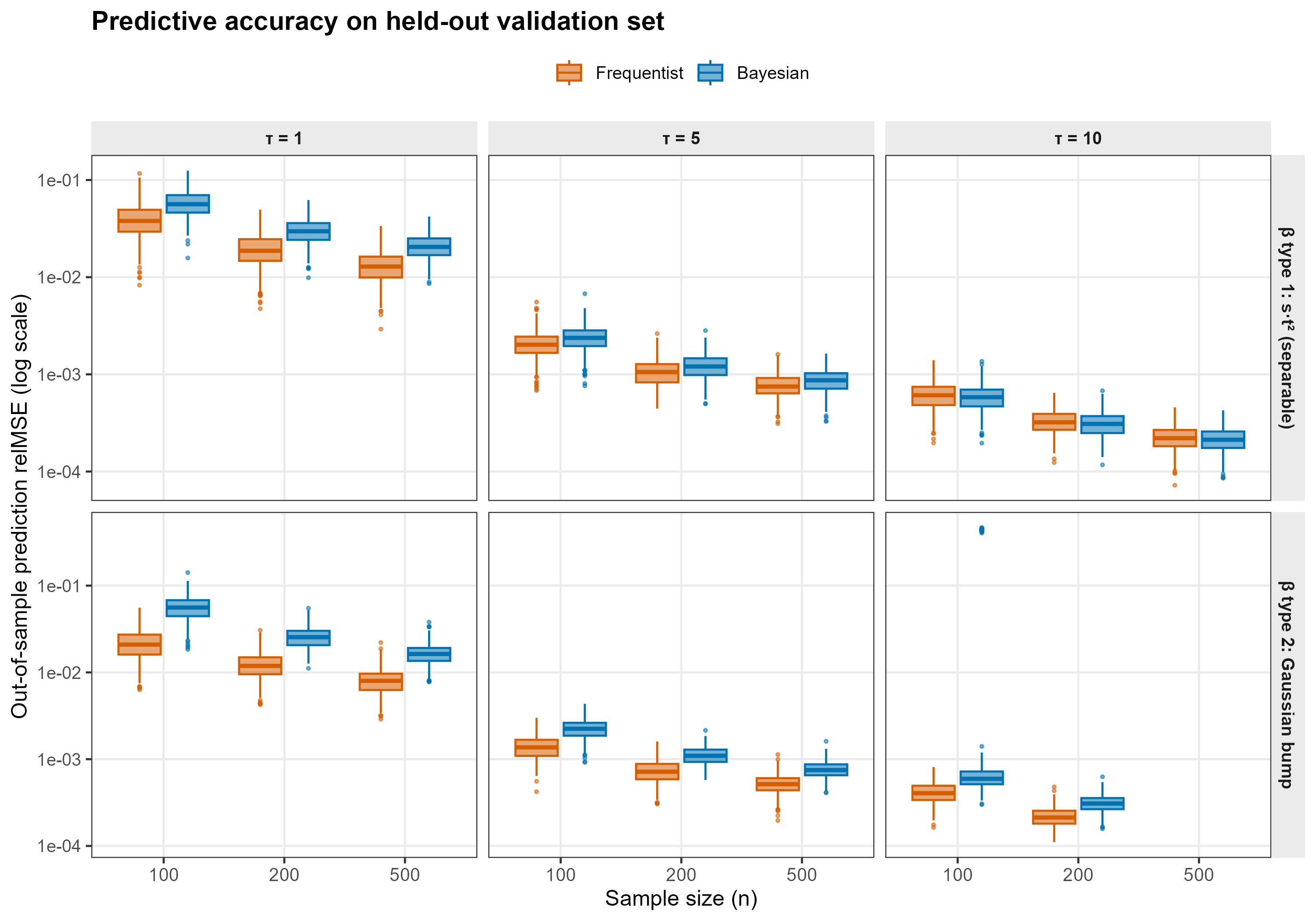
Performance Metric
For each replicate we draw an independent validation set of subjects from the same data-generating process, evaluate each method’s predicted mean response , and compute the relative prediction MSE
This metric is invariant to the unidentifiable
additive-in-
component of
(see the package’s Simulation/FoFR_identifiability_note.md
for details), so it provides an apples-to-apples comparison even though
pffr() includes an explicit functional intercept and
fofr_bayes() does not.
References
- Jiang, Z., Crainiceanu, C., and Cui, E. (2025). Tutorial on Bayesian Functional Regression Using Stan. Statistics in Medicine, 44(20–22), e70265.
- Crainiceanu, C. M., Goldsmith, J., Leroux, A., and Cui, E. (2024). Functional Data Analysis with R. CRC Press.
- Goldsmith, J., Zipunnikov, V., and Schrack, J. (2015). Generalized Multilevel Function-on-Scalar Regression and Principal Component Analysis. Biometrics, 71(2), 344–353.
- Ramsay, J. O. and Silverman, B. W. (2005). Functional Data Analysis, 2nd Edition. Springer.
- Ivanescu, A. E., Staicu, A.-M., Scheipl, F., and Greven, S. (2015). Penalized Function-on-Function Regression. Computational Statistics, 30(2), 539–568.
- Scheipl, F., Staicu, A.-M., and Greven, S. (2015). Functional Additive Mixed Models. Journal of Computational and Graphical Statistics, 24(2), 477–501.
- Carpenter, B., Gelman, A., Hoffman, M. D., et al. (2017). Stan: A Probabilistic Programming Language. Journal of Statistical Software, 76(1), 1–32.